-
Metagenome
-
Metagenome analysis is an analysis that identifies and classifies various bacteria, viruses, and fungi that exist in the organs (intestinal, respiratory, skin, and etc.) of living organisms or in environmental units such as the sea and soil.
-
Metagenomic NGS
-
PHYZEN provides species identification, network analysis, diversity analysis through amplicon sequencing using the Gold Standard regions like bacterial 16S rRNA genes and fungal ITS regions as well as functional profiling in microbial communities using whole-genome and whole-transcriptome sequencing.
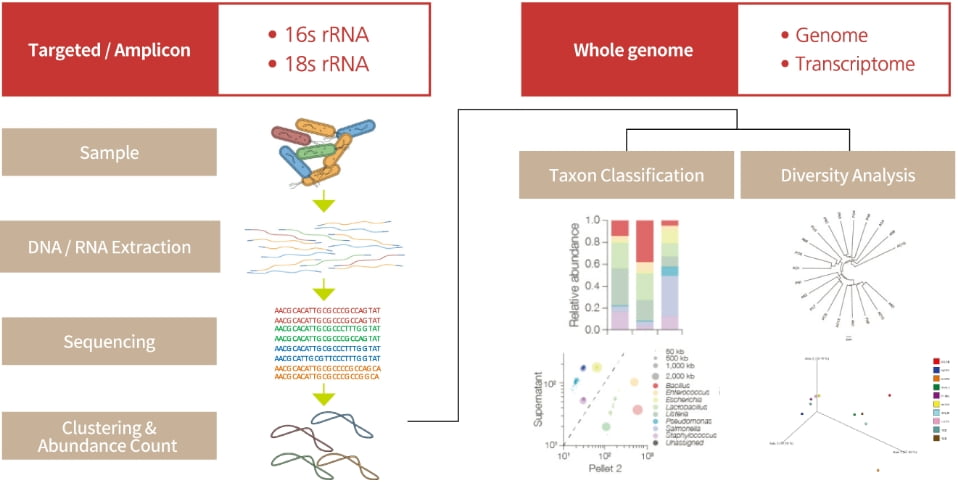
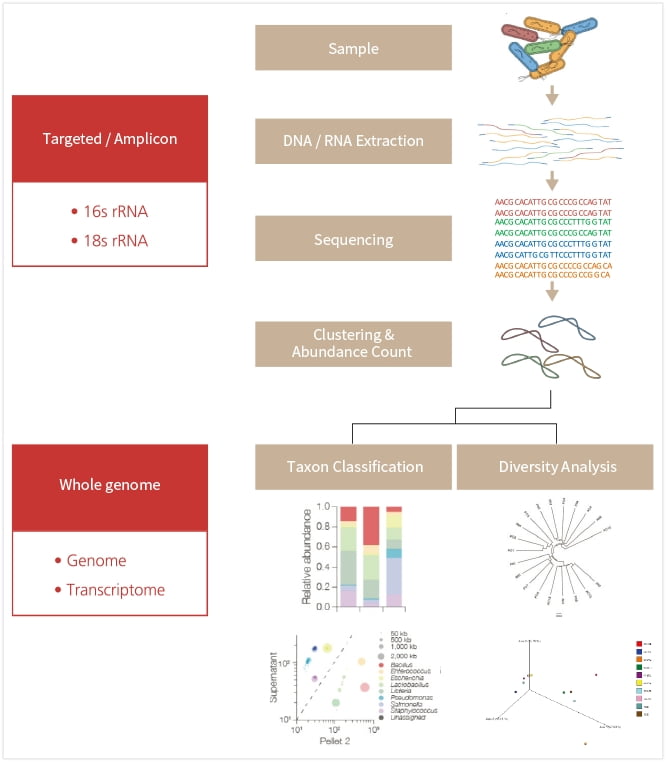
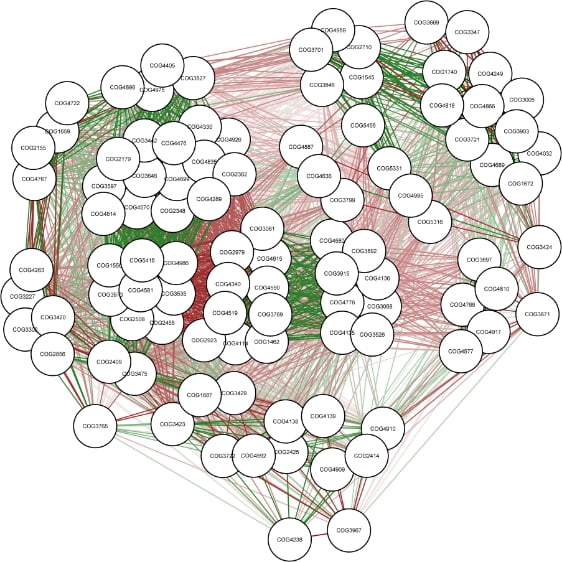
Network analysis among microbial species
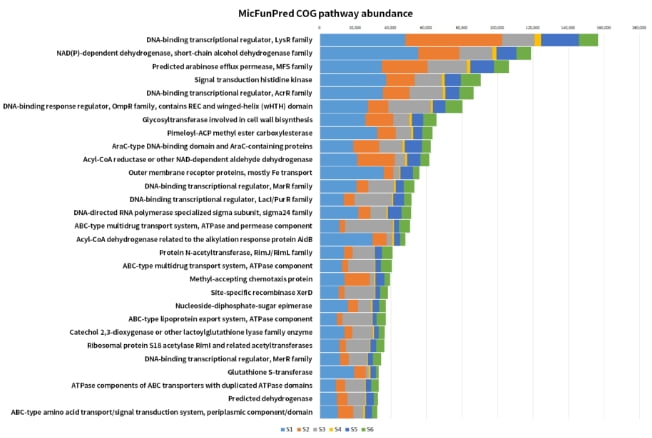
Functional profiling
-
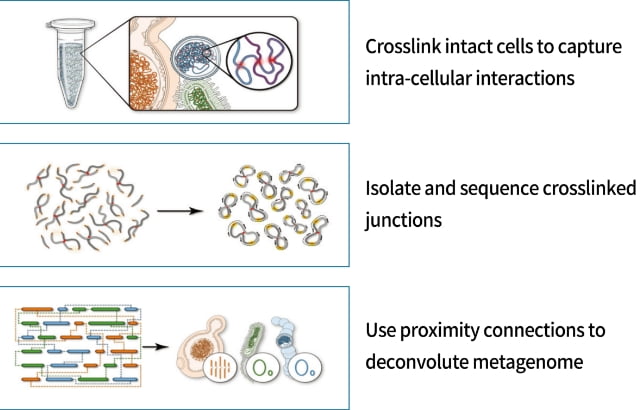
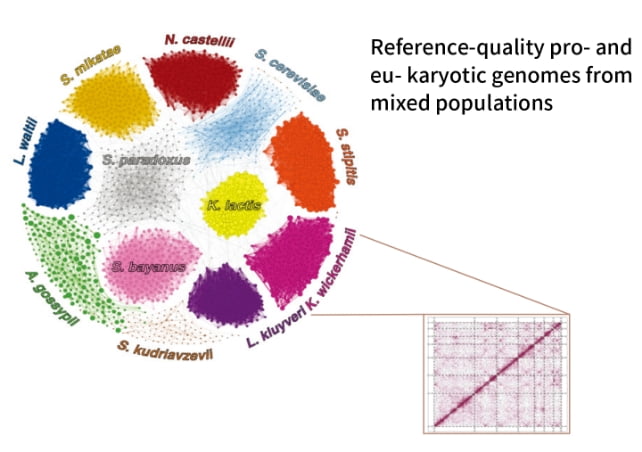
Metagenomic Hi-C
-
Simultaneous multiple genome assembly and classification of microbial communities are possible using whole genome metagenome analysis combined with Hi-C analysis.